

| Pathway Id | ko03015 |
| Class & Sub-Class | Class: Genetic Information Processing Sub-Class: Translation |
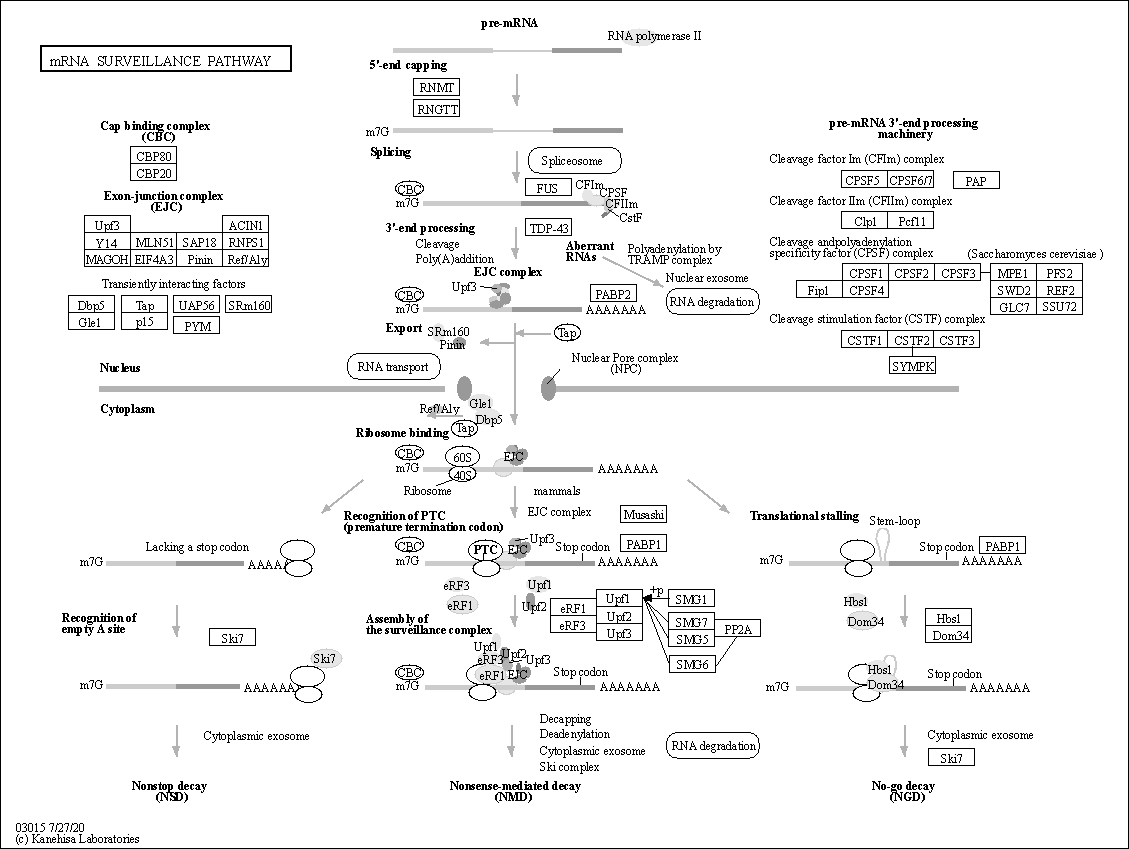
| Description | The mRNA surveillance pathway is a quality control mechanism that detects and degrades abnormal mRNAs. These pathways include nonsense-mediated mRNA decay (NMD); nonstop mRNA decay (NSD); and no-go decay (NGD). NMD is a mechanism that eliminates mRNAs containing premature translation-termination codons (PTCs). In vertebrates; PTCs trigger efficient NMD when located upstream of an exon junction complex (EJC). Upf3; together with Upf1 and Upf2; may signal the presence of the PTC to the 5'end of the transcript; resulting in decapping and rapid exonucleolytic digestion of the mRNA. In the NSD pathway; which targets mRNAs lacking termination codons; the ribosome is believed to translate through the 3' untranslated region and stall at the end of the poly(A) tail. NSD involves an eRF3-like protein; Ski7p; which is hypothesized to bind the empty A site of the ribosome and recruit the exosome to degrade the mRNA from the 3' end. NGD targets mRNAs with stalls in translation elongation for endonucleolytic cleavage in a process involving the Dom34 and Hbs1 proteins. Source: KEGG Database |
Make changes to the option below (image Size) and click here to update the map:
| Image Size | You can change the size of an image just by changing the width or height attributes. Size (enter width only) the present size is: | |
| Get Data Out | |
| Click here to show selected items: | |